Background
Small RNA sequencing (RNA-Seq) is a technique to isolate and sequence small RNA species, such as microRNAs (miRNAs), short interfering RNA (siRNA), piwi-interacting RNA (piRNA), and more. Small RNA-Seq can query thousands of small RNA and miRNA sequences with unprecedented sensitivity and dynamic range. With small RNA-Seq you can discover novel miRNAs and other small noncoding RNAs, and examine the differential expression of all small RNAs in any sample.
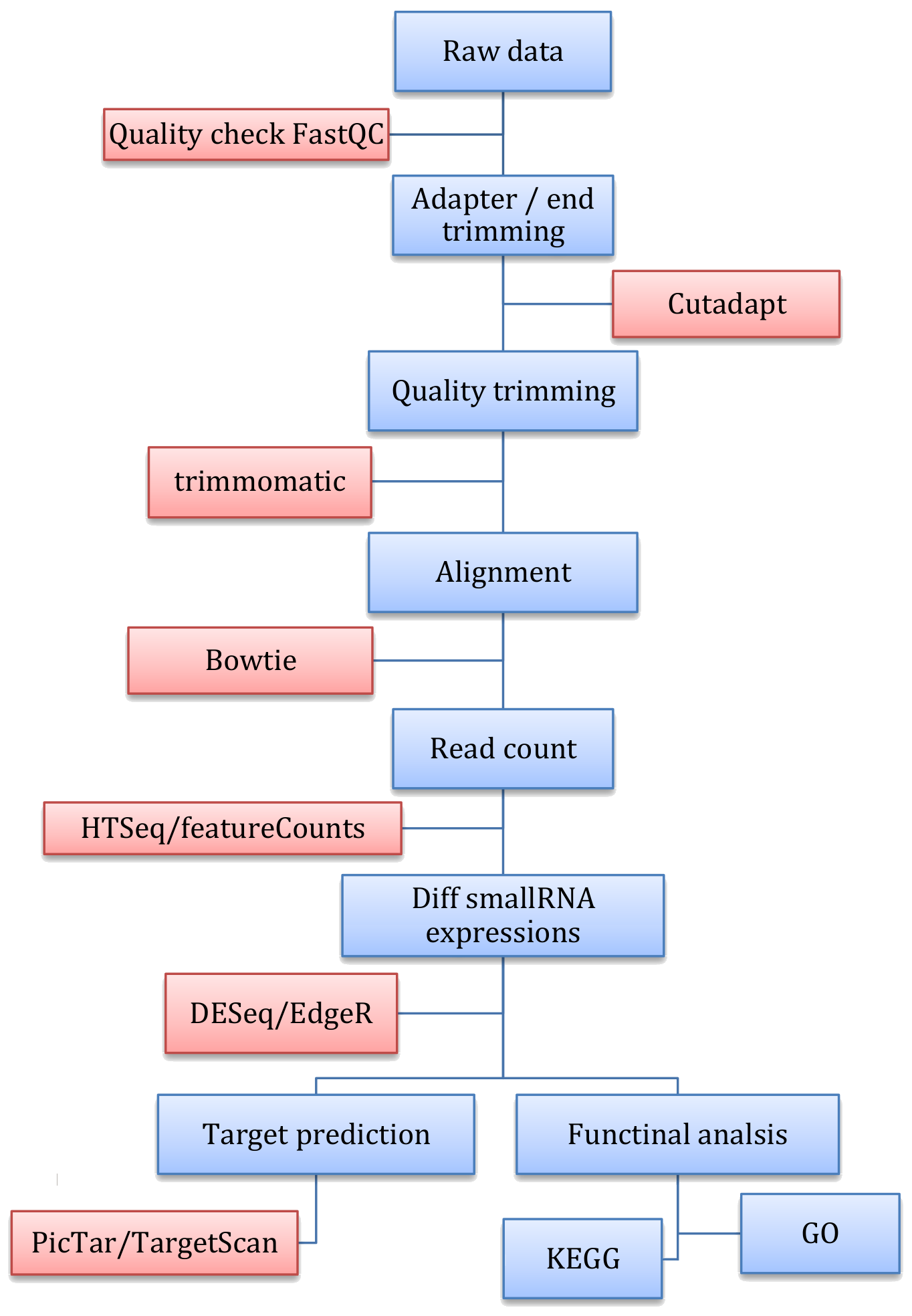
Figure: smallRNA-Seq workflow